TLDR; You can use the corrtable package (see CRAN or Github)!
In most (observational) research papers you read, you will probably run into a correlation matrix. Often it looks something like this:

In Social Sciences, like Psychology, researchers like to denote the statistical significance levels of the correlation coefficients, often using asterisks (i.e., *). Then the table will look more like this:

Regardless of my personal preferences and opinions, I had to make many of these tables for the scientific (non-)publications of my Ph.D..
I remember that, when I first started using R, I found it quite difficult to generate these correlation matrices automatically.
Yes, there is the cor function, but it does not include significance levels.
Then there the (in)famous Hmisc package, with its rcorr function. But this tool provides a whole new range of issues.
What’s this storage.mode, and what are we trying to coerce again?
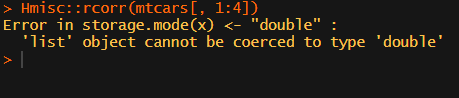
Soon you figure out that Hmisc::rcorr only takes in matrices (thus with only numeric values). Hurray, now you can run a correlation analysis on your dataframe, you think…
Yet, the output is all but publication-ready!
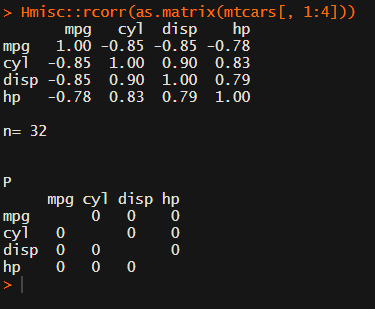
You wanted one correlation matrix, but now you have two… Double the trouble?
[UPDATED] To spare future scholars the struggle of the early day R programming, Laura Lambert and I created an R package corrtable, which includes the helpful function correlation_matrix.
This correlation_matrix takes in a dataframe, selects only the numeric (and boolean/logical) columns, calculates the correlation coefficients and p-values, and outputs a fully formatted publication-ready correlation matrix!
You can specify many formatting options in correlation_matrix.
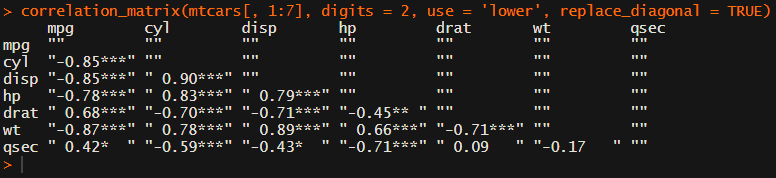
For instance, you can use only 2 decimals. You can focus on the lower triangle (as the lower and upper triangle values are identical). And you can drop the diagonal values:
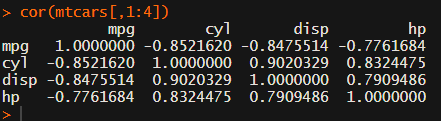
Or maybe you are interested in a different type of correlation coefficients, and not so much in significance levels:
For other formatting options, do have a look at the source code on github.
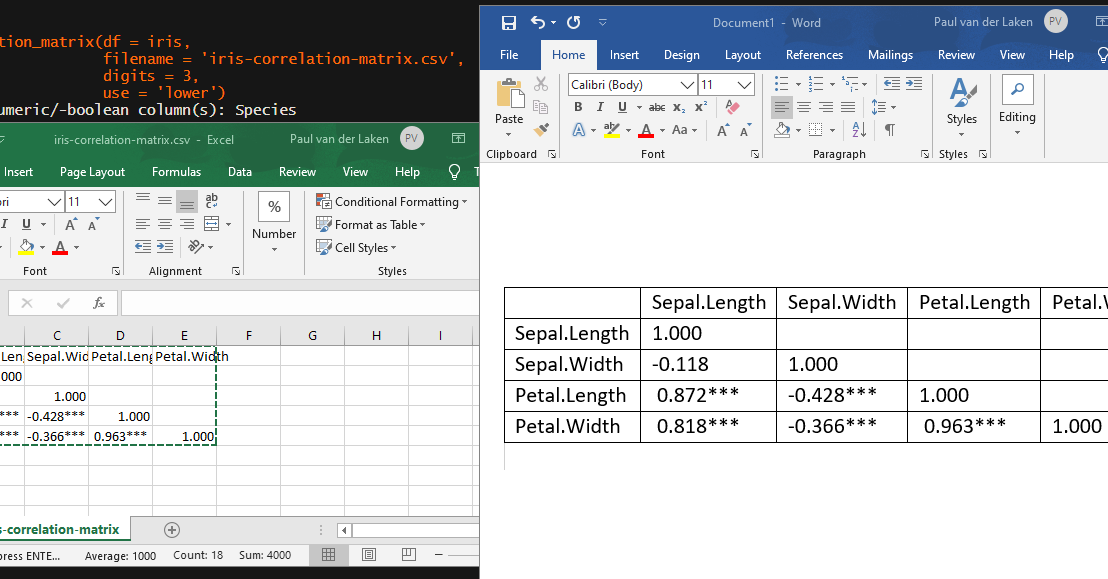
Now, to make matters even easier, the package includes a second function (save_correlation_matrix) to directly save any created correlation matrices:
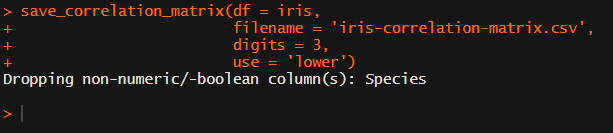
Once you open your new correlation matrix file in Excel, it is immediately ready to be copy-pasted into Word!

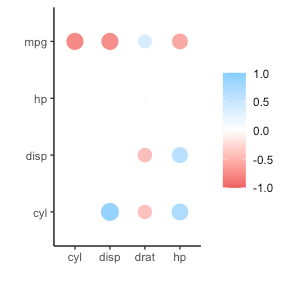
If you are looking for ways to visualize your correlations do have a look at the packages corrr, corrplot, or ppsr.
I hope this package is of help to you!
Do reach out if you get to use them in any of your research papers!
Sign up to keep up to date on the latest R, Data Science & Tech content: